Subsetting and Filtering a Data Frame in R (base R)
When I was first learning R in a Coursera course from Johns Hopkins University, subsetting and filtering was one of the first things I learned how to do in R. Subsetting is essentially scaling down your data frame so that you are only seeing relevant data points. Filtering a data frame is super important to know how to do, since data frames, in my opinion, are the most common data structures you’ll use in R.
While I think it is extremely important for those learning R to have a good foundation in base R code, I know that there are several packages out there that make subsetting and filtering data frames easier and faster. We’ll get in to those later, but for now, let’s look at the base R way of doing things.
Subsetting the Base R Way
When I say Base R, I am referring to using straight R code and not introducing any additional packages to be loaded.
The general format for subsetting a data frame looks like this:
dataframe[ row, column ]
The row and column parameters between the brackets can be a single index, a range (1:10), a character vector containing multiple indexes (c(1,3,5,7,9)), or left blank to return all rows. The column parameter can take the same options, and it will also accept column names. Subsetting allows you to scale down the dataset with which you are working.
A best practice that I use is to assign the subset to a new variable so that the original information is not lost.
newdataframe = dataframe[1:100, c("col1", "col2", "col6")]
When I was first learning to subset data frames in R, I preferred to use the column indexes because I didn’t need to have opening and closing quotes, saving me time. I changed my ways a long time ago when I got bit by an automated report that was modified. The column names were not in the same order anymore and tons of errors were in the console.
You’ll thank me later: If at all possible, reference the columns by column name to save yourself frustration and refactoring in the long run.
Subsetting Examples
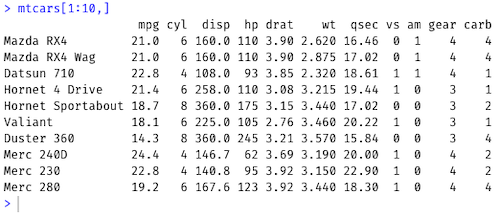
Show the first 10 rows of the mtcars dataset and all columns:
mtcars[1:10, ]
Show the first 3 rows, but only see the mpg and cyl columns:
mtcars[1:3, c(1,2)] or mtcars[1:3, c("mpg", "cyl")]
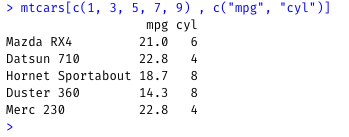
Show rows 1, 3, 5, 7 and 9, and only the values from the mpg and cyl columns:
mtcars[c(1, 3, 5, 7, 9) , c("mpg", "cyl")]
Filtering the Base R Way
If you want to filter a data frame, you’ll add the logic to the row parameter in the brackets. This is where it can get confusing to write R code using base R.
To filter a data frame based on a column, you’ll use the following format:
dataframe[ dataframe$column >= 21, column ].
The >= 21 part is where you’ll add your conditional logic for the filter. Additional logic can be used by adding the and operator (&) or the or operator (|). It always seemed backwards in my head to add the logic to the rows and not the columns, but eventually it clicked, and off I went.
The column parameter functions identically to how it does when subsetting a data frame. The column parameter will accept a single index, a range (1:10), a character vector containing multiple indexes or column names in quotes, or left blank to return all columns.
Filtering Examples
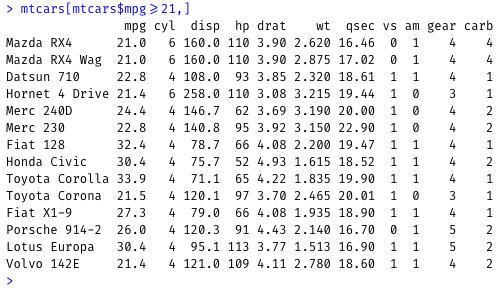
Filter the mtcars dataset to show cars that mpg values greater than or equal to 21:
mtcars[mtcars$mpg>=21,]
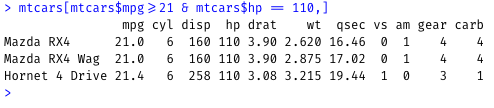
Filter the mtcars dataset to show cars that mpg values greater than or equal to 21 and have horsepower (hp) of 100:
mtcars[mtcars$mpg>=21 & mtcars$hp == 110,]
Show cars that have 3 or 5 gears:
mtcars[mtcars$gear == 3 | mtcars$gear == 5, ]
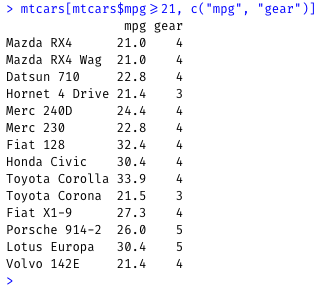
Show cars that get 21 or more mpg, and only show the mpg and gear columns:
mtcars[mtcars$mpg>=21, c("mpg", "gear")]
Conclusion
Subsetting and filtering data frames in R using the base R code is super important on your coding journey. It’s best to learn the base R way of doing things so that down the road, you’ll be able to troubleshoot errors and understand why there are conflicts with packages. That said, subsetting and filtering in R can be done faster and easier (in my opinion) with the help of the dplyr package. This package is very powerful for manipulating data frames, and it is extremely well documented.